biomod2 team videos
-
All first steps of your biomod2 modeling, going from formatting
of your data, to defining of your modeling options, through
pseudo-absences and cross-validation data sets.
- All information about single models within biomod2, introducing data types and algorithms, and exploring evaluation metrics, variables’ importance and response curves.
Chapters can be directly accessed to through time stamps in the video description on YouTube.
Tutorial 2 - v4.3-4 - part 1
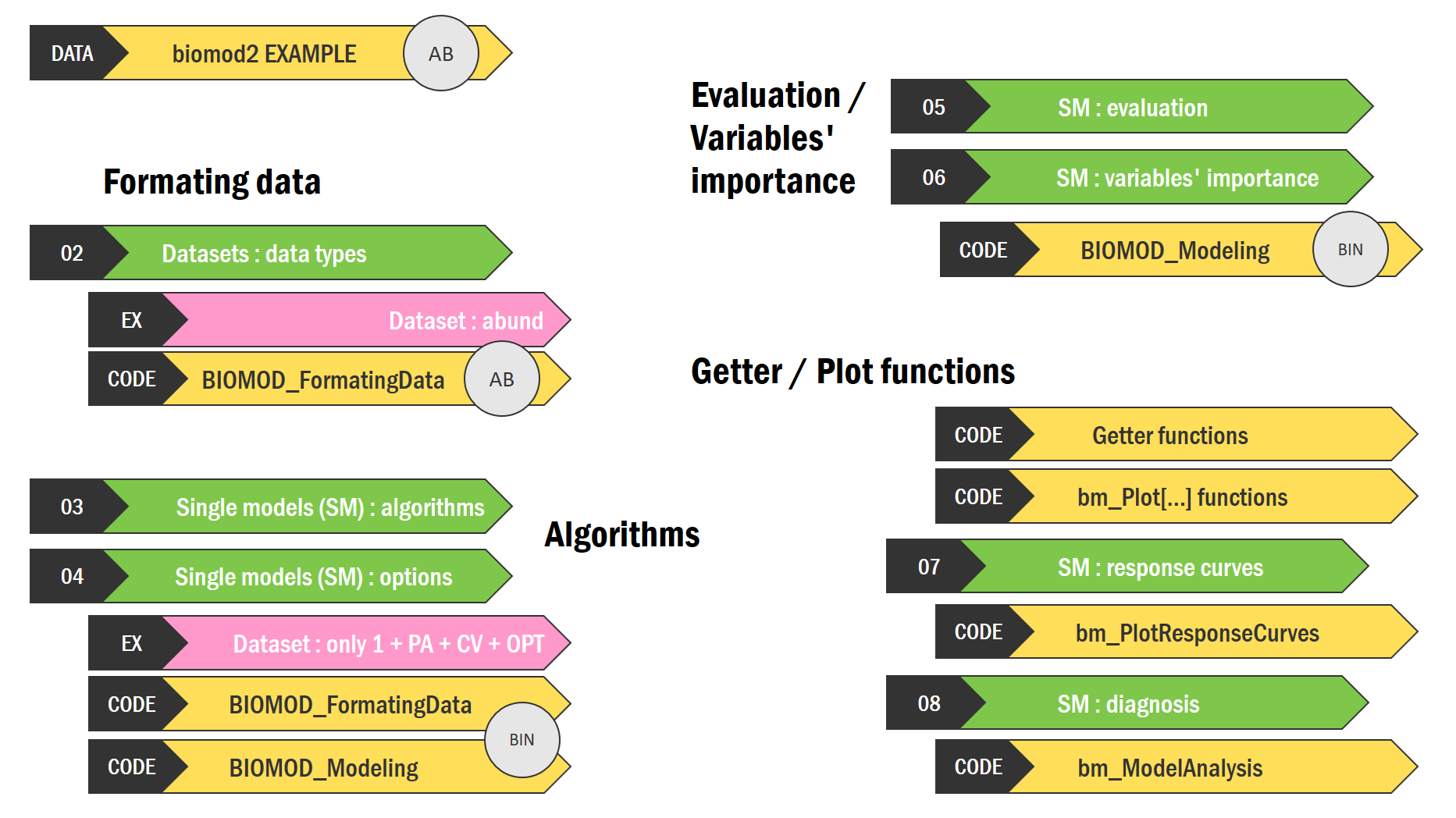
| Chapter | Theme | Timestamp |
|---|---|---|
| Evolution through time | 0:00 | |
| 01. Functions hierarchy | 1:00 | |
| DATA. biomod2 example (BIN / AB) | 1:32 | |
| FORMATING DATA | 2:33 | |
| 02. Datasets : data types | 3:19 | |
| Dataset : abund | 4:38 | |
| ALGORITHMS | 5:49 | |
| 03. Single models : algorithms | 6:51 | |
| 04. Single models : options | 9:00 | |
| Dataset : only 1 + PA + CV + OPT (BIN) | 10:28 | |
| EVALUATION / VARIABLES’ IMPORTANCE | 05. Single models : evaluation | 15:10 |
| 06. Single models : variables’ importance | 19:38 | |
| Code : BIOMOD_Modeling(BIN) | 22:38 | |
| GETTER / PLOT FUNCTIONS | 23:41 | |
| Code : Getter functions | 24:49 | |
| Code : Plot functions | 29:09 | |
| 07. Single models : response curves | 32:47 | |
| 08. Single models : diagnosis | 36:43 |
Tutorial 1 - v4.2-6
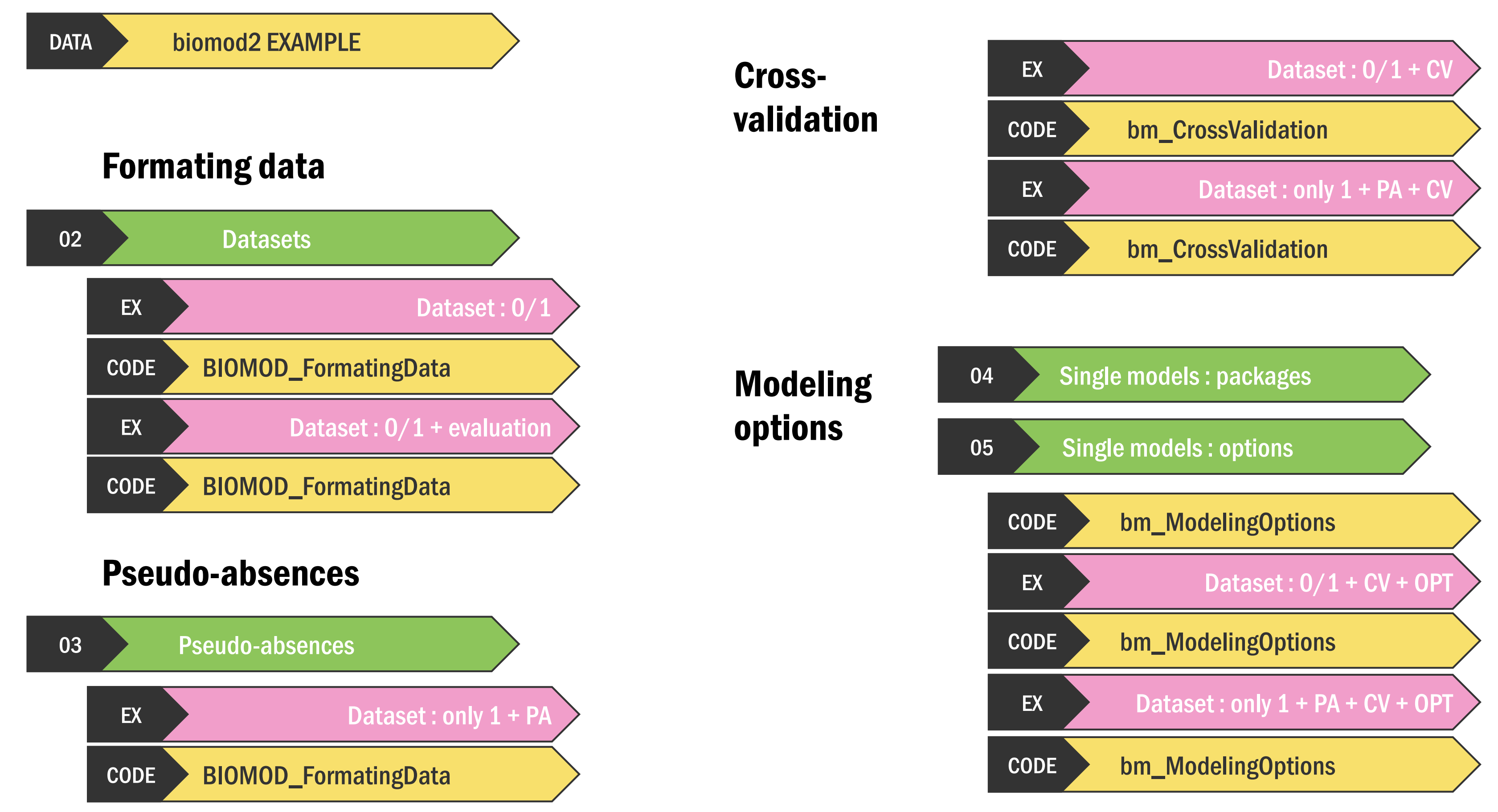
| Chapter | Theme | Timestamp |
|---|---|---|
| Introduction to the package | 0:00 | |
| 01. Functions hierarchy | 2:28 | |
| DATA. biomod2 example | 3:54 | |
| FORMATING DATA | 02. Datasets | 4:14 |
| Dataset : 0/1 | 7:07 | |
| Dataset : 0/1 + evaluation | 8:21 | |
| PSEUDO-ABSENCES | 9:08 | |
| 03. Pseudo-absences | 9:39 | |
| Dataset : only 1 + PA | 13:05 | |
| CROSS-VALIDATION | Dataset : 0/1 + CV | 16:35 |
| Dataset : only 1 + PA + CV | 18:39 | |
| MODELING OPTIONS | 04. Single models : packages | 19:49 |
| 05. Single models : options | 20:26 | |
| Dataset : 0/1 + CV + OPT | 24:48 | |
| Dataset : only 1 + PA + CV + OPT | 26:40 |